Multi-Dimensional Single Cell Measurement Technologies

This artistic image represents microfluidic antigen-TCR engagement sequencing, which was highlighted in an article published in the journal Lab on a Chip. This image, created at ISB by Alex Xu, Alphonsus Ng, Allison Kudla and Stella Sun, was featured as cover art by the publication.
A challenge associated with single cell omics methods is to capture quantitative, multiplex information from single cells, while retaining the tissue context of those cells. New programs in the Heath group are focused on addressing this challenge by developing tools to capture information from distinct areas of biology in the same single cells: to determine how cellular energetics relates to gene expression, or protein function, or cell-cell binding.
Our goal is to apply these tools to better understand the systems biology of disease.We develop and engineer an array of methods to study single cells, including the single cell barcode chip (SCBC) and customized metabolic probes, developed in the Heath lab; Nanostring digital spatial profiling, an ISB spin-off; and single cell RNA sequencing tools like MATE-seq. Our technology feeds into bioinformatic inference tools developed by collaborators, which help us discover connections across the spectrum of biology.
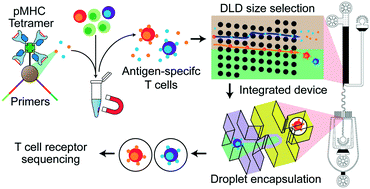
Microfluidic antigen-TCR engagement sequencing (MATE-seq)
Featured Publication
Integrated measurement of intracellular proteins and transcripts in single cells, Lab on a Chip (2018). Go to →
Surface Immobilization of Redox‐Labile Fluorescent Probes: Enabling Single‐Cell Co‐Profiling of Aerobic Glycolysis and Oncogenic Protein Signaling Activities, Angewandte Chemie International Edition (2018). Go to →
MATE-Seq: microfluidic antigen-TCR engagement sequencing, Lab on a Chip (2019). Go to→